sample_arc_from_normal()
Sample mesh data along a circular arc defined by a normal vector.
Creates a circular arc probe defined by a normal to the plane of the arc, a polar starting vector, and an angle. The arc is sampled in a counterclockwise direction from the polar vector. For temporal datasets, automatically sweeps all time points unless time_value is specified.
Parameters
center(Sequence[float]) — Center point[x, y, z]of the circle that defines the arc.resolution(int, default:100) — Number of segments to divide the arc into. The resulting arc will haveresolution + 1points.normal(Sequence[float] | None, default:None) — Normal vector[x, y, z]to the plane of the arc. IfNone, defaults to[0, 0, 1](positive Z direction).polar(Sequence[float] | None, default:None) — Starting point of the arc in polar coordinates[x, y, z]. IfNone, defaults to[1, 0, 0](positive X direction).angle(float | None, default:None) — Arc length in degrees, beginning at the polar vector in a counterclockwise direction. IfNone, defaults to 90 degrees.time_value(float | None, default:None) — Query a specific time value instead of sweeping all time points. Ignored for static datasets.tolerance(float | None, default:None) — Tolerance for the sample operation. IfNone, PyVista generates a tolerance automatically.
Returns
Returns a list of dictionaries (one per time step) with a "time" key and array names as keys. Each array has:
- For scalars:
"distance","value","x","y","z"(sample point coordinates) - For vectors:
"distance","x_value","y_value","z_value","tangential"(component along arc),"normal"(component perpendicular to arc),"x","y","z"(sample point coordinates)
For static datasets, returns a single-element list with time 0.0.
[
# Time 0 (or single time for static)
{
"time": 0.0,
"scalar_name": {
"distance": [0.0, 0.1, 0.2, ...],
"value": [val0, val1, val2, ...],
"x": [x0, x1, x2, ...],
"y": [y0, y1, y2, ...],
"z": [z0, z1, z2, ...]
},
"vector_name": {
"distance": [0.0, 0.1, 0.2, ...],
"x_value": [x0, x1, x2, ...],
"y_value": [y0, y1, y2, ...],
"z_value": [z0, z1, z2, ...],
"tangential": [t0, t1, t2, ...],
"normal": [n0, n1, n2, ...],
"x": [x0, x1, x2, ...],
"y": [y0, y1, y2, ...],
"z": [z0, z1, z2, ...]
},
...
},
# Time 1
{"time": 0.01, ...},
...
]
Examples
Sweep all time steps
from pyemsi import Plotter, examples
from matplotlib import pyplot
import numpy as np
file_path = examples.ipm_motor_path()
plt = Plotter(file_path)
data = plt.sample_arc_from_normal(
center=(0, 0, 0),
normal=(0, 0, 1),
polar=(0.080575, 0, 0),
angle=45,
resolution=100,
)
time_values = [time_data["time"] for time_data in data]
distances = data[0]["B-Mag (T)"]["distance"]
value_grid = np.array([time_data["B-Mag (T)"]["value"] for time_data in data])
time_grid, distance_grid = np.meshgrid(time_values, distances, indexing="ij")
fig, ax = pyplot.subplots(subplot_kw={"projection": "3d"})
ax.plot_surface(time_grid, distance_grid, value_grid, cmap="viridis")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Distance Along Arc (m)")
ax.set_zlabel("B-Mag (T)")
ax.set_title("B-Mag (T) Along Sampled Arc")
fig.tight_layout()
fig.savefig("docs/static/demos/sample_arc_from_normal.png")
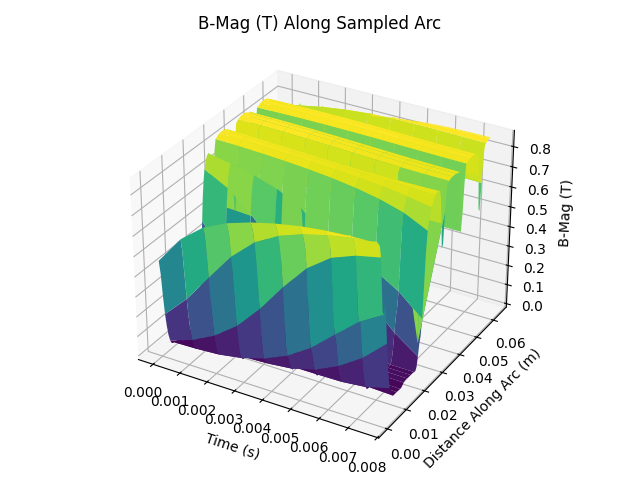
Plot three time slices
from pyemsi import Plotter, examples
from matplotlib import pyplot
file_path = examples.ipm_motor_path()
plt = Plotter(file_path)
data = plt.sample_arc_from_normal(
center=(0, 0, 0),
normal=(0, 0, 1),
polar=(0.080575, 0, 0),
angle=45,
resolution=100,
)
time_indices = sorted({0, len(data) // 2, len(data) - 1})
fig, ax = pyplot.subplots()
for idx in time_indices:
ax.plot(
data[idx]["B-Mag (T)"]["distance"],
data[idx]["B-Mag (T)"]["value"],
label=f"t = {data[idx]['time']:.3f} s",
)
ax.set_xlabel("Distance Along Arc (m)")
ax.set_ylabel("B-Mag (T)")
ax.set_title("B-Mag (T) Along Sampled Arc at Three Time Points")
ax.legend()
fig.tight_layout()
fig.savefig("docs/static/demos/sample_arc_from_normal_time_slices.png")
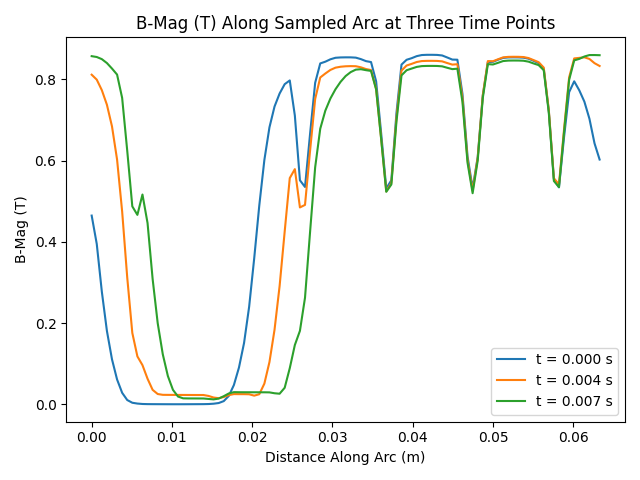
Plot tangential and normal B-Vec components
from pyemsi import Plotter, examples
from matplotlib import pyplot
import numpy as np
file_path = examples.ipm_motor_path()
plt = Plotter(file_path)
data = plt.sample_arc_from_normal(
center=(0, 0, 0),
normal=(0, 0, 1),
polar=(0.080575, 0, 0),
angle=45,
resolution=100,
)
time_indices = sorted({0, len(data) // 2, len(data) - 1})
fig, axes = pyplot.subplots(1, 2, figsize=(12, 4))
component_map = [
("tangential", "Tangential B-Vec (T)"),
("normal", "Normal B-Vec (T)"),
]
for ax, (component_key, ylabel) in zip(np.atleast_1d(axes), component_map):
for idx in time_indices:
ax.plot(
data[idx]["B-Vec (T)"]["distance"],
data[idx]["B-Vec (T)"][component_key],
label=f"t = {data[idx]['time']:.3f} s",
)
ax.set_xlabel("Distance Along Arc (m)")
ax.set_ylabel(ylabel)
ax.legend()
axes[0].set_title("Tangential Component")
axes[1].set_title("Normal Component")
fig.suptitle("B-Vec (T) Components Along Sampled Arc at Three Time Points")
fig.tight_layout()
fig.savefig("docs/static/demos/sample_arc_from_normal_bvec_components_time_slices.png")
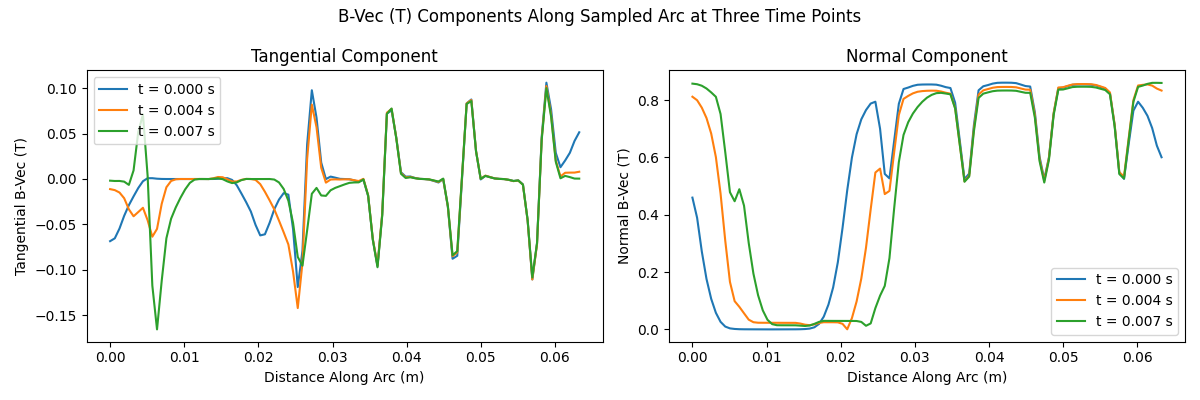
Notes
- The arc is constructed in a counterclockwise direction from the polar vector as viewed from the direction of the normal vector.
- The default normal
[0, 0, 1]and polar[1, 0, 0]create an arc in the XY plane starting from the positive X axis. - For vectors,
tangentialrepresents the component along the arc direction, whilenormalrepresents the magnitude perpendicular to the arc. - The
"x","y","z"keys contain the actual 3D coordinates of sample points along the arc. - This method is useful when you want to define an arc by plane orientation rather than explicit start/end points.
See Also
sample_arc()- Sample along arc defined by start/end pointssample_arcs_from_normal()- Sample multiple arcs defined by normal vectorssample_line()- Sample along straight line