sample_lines()
Sample mesh data along multiple straight lines.
Creates line probes for each (pointa, pointb) pair and samples the mesh data onto each line. For temporal datasets, automatically sweeps all time points unless time_value is specified.
Parameters
lines(Sequence[tuple[Sequence[float], Sequence[float]]]) — List of line definitions, each as a tuple(pointa, pointb)wherepointaandpointbare[x, y, z]coordinates.resolution(int | list[int], default:100) — Number of segments to divide each line into. Can be a singleint(applied to all lines) or a list ofints (one per line).time_value(float | None, default:None) — Query a specific time value instead of sweeping all time points. Ignored for static datasets.tolerance(float | None, default:None) — Tolerance for the sample operation. IfNone, PyVista generates a tolerance automatically.progress_callback(callable | None, default:None) — Callback function for progress updates. Called with(current_line, total_lines). Should returnTrueto continue orFalseto cancel.
Returns
Returns a list of results (one per line), where each result is a list of dictionaries (one per time step) with a "time" key and array names as keys. Each array has:
- For scalars:
"distance","value","x","y","z"(sample point coordinates) - For vectors:
"distance","x_value","y_value","z_value","tangential","normal","x","y","z"(sample point coordinates)
[
# First line results
[
# Time 0
{"time": 0.0, "scalar_name": {"distance": [...], "value": [...], "x": [...], "y": [...], "z": [...]}, ...},
# Time 1
{"time": 0.01, "scalar_name": {"distance": [...], "value": [...], "x": [...], "y": [...], "z": [...]}, ...},
...
],
# Second line results
[
{"time": 0.0, "scalar_name": {"distance": [...], "value": [...], "x": [...], "y": [...], "z": [...]}, ...},
...
],
...
]
Examples
Sweep all time steps
from pyemsi import Plotter, examples
from matplotlib import pyplot
import numpy as np
file_path = examples.transient_path()
plt = Plotter(file_path)
data = plt.sample_lines(
lines=[
((0.0, 0.0, 0.0), (0.0, 0.0, 0.25)),
((0.02, 0.02, 0.0), (0.02, 0.02, 0.25)),
],
resolution=100,
)
fig, axes = pyplot.subplots(1, len(data), figsize=(14, 6), subplot_kw={"projection": "3d"})
axes = np.atleast_1d(axes)
for idx, line_data in enumerate(data):
time_values = [time_data["time"] for time_data in line_data]
distances = line_data[0]["B-Mag (T)"]["distance"]
value_grid = np.array([time_data["B-Mag (T)"]["value"] for time_data in line_data])
time_grid, distance_grid = np.meshgrid(time_values, distances, indexing="ij")
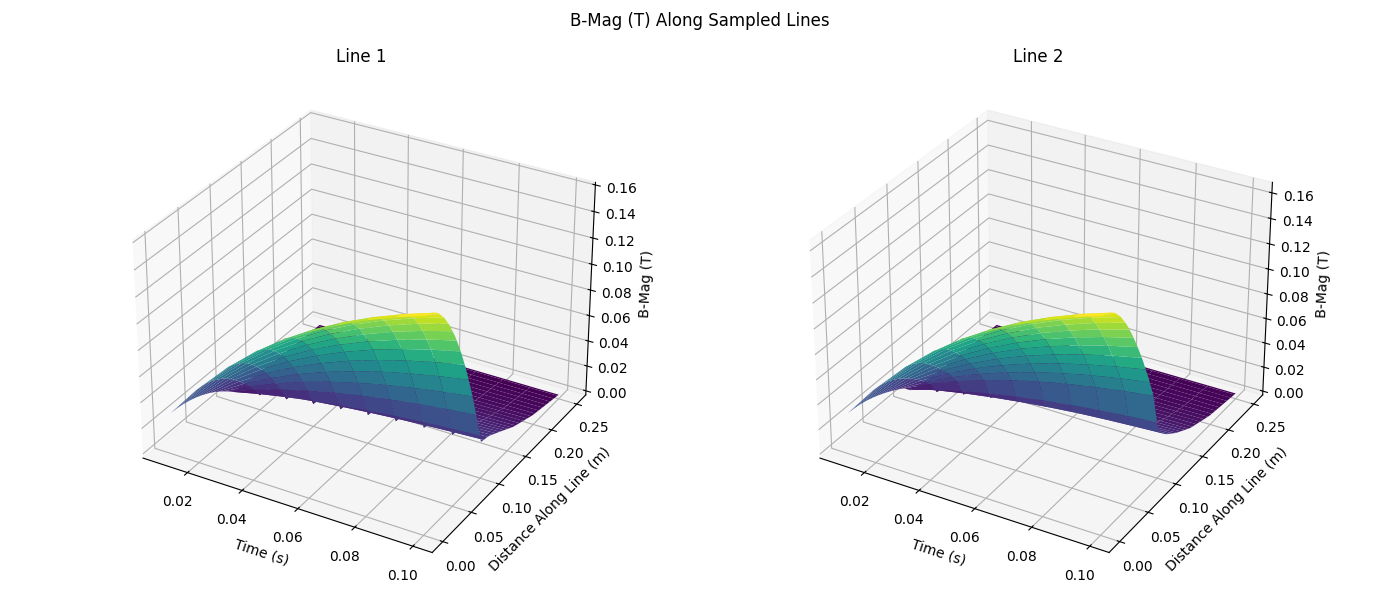
axes[idx].plot_surface(time_grid, distance_grid, value_grid, cmap="viridis")
axes[idx].set_xlabel("Time (s)")
axes[idx].set_ylabel("Distance Along Line (m)")
axes[idx].set_zlabel("B-Mag (T)")
axes[idx].set_title(f"Line {idx + 1}")
fig.suptitle("B-Mag (T) Along Sampled Lines")
fig.tight_layout()
fig.savefig("docs/static/demos/sample_lines.png")
Plot three time slices
from pyemsi import Plotter, examples
from matplotlib import pyplot
import numpy as np
file_path = examples.transient_path()
plt = Plotter(file_path)
data = plt.sample_lines(
lines=[
((0.0, 0.0, 0.0), (0.0, 0.0, 0.25)),
((0.02, 0.02, 0.0), (0.02, 0.02, 0.25)),
],
resolution=100,
)
fig, axes = pyplot.subplots(1, len(data), figsize=(14, 5))
axes = np.atleast_1d(axes)
for idx, line_data in enumerate(data):
time_indices = sorted({0, len(line_data) // 2, len(line_data) - 1})
for time_idx in time_indices:
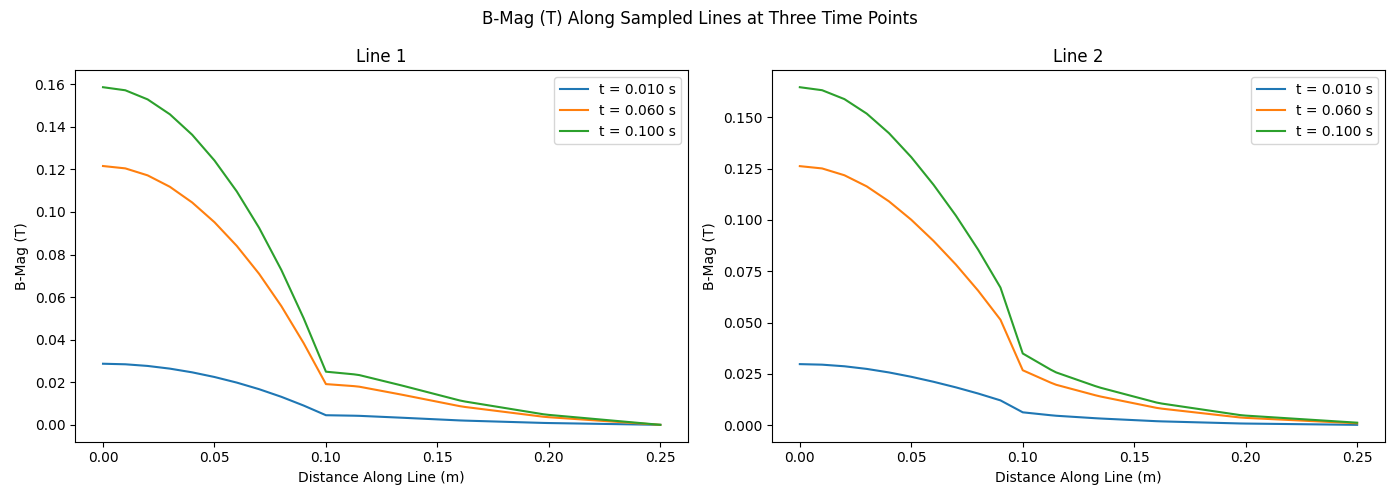
axes[idx].plot(
line_data[time_idx]["B-Mag (T)"]["distance"],
line_data[time_idx]["B-Mag (T)"]["value"],
label=f"t = {line_data[time_idx]['time']:.3f} s",
)
axes[idx].set_xlabel("Distance Along Line (m)")
axes[idx].set_ylabel("B-Mag (T)")
axes[idx].set_title(f"Line {idx + 1}")
axes[idx].legend()
fig.suptitle("B-Mag (T) Along Sampled Lines at Three Time Points")
fig.tight_layout()
fig.savefig("docs/static/demos/sample_lines_time_slices.png")
See Also
sample_line()— Sample along a single linesample_arcs()— Sample along multiple circular arcssample_points()— Sample at multiple point coordinates